Scientists were in constant search for efficient and precise genome editing using reagents that are simple to design as well as inexpensive to generate. Developing ways to make precise, targeted changes to the genome of living cells has been a long-standing goal of biomedical researchers, who seek to understand, and hopefully cure, genetic diseases like sickle-cell anemia and cystic fibrosis.
Genome editing is a type of genetic engineering that uses artificially engineered nucleases, or “molecular scissors,” to insert, replace, or remove DNA from a genome. In recent years several methods have been developed that are revolutionizing the field. All allow for a relatively quick and targeted change of one or several genes in any given genome. The targeted genes are either turned off or changed by additionally adding engineered pieces of DNA.

Index
Different types of Nucleases
Nucleases can be designed to bind to specific DNA target sequences and create precise double-strand breaks.
Meganucleases
Meganucleases are the first target nucleases developed in the year 2003. These are large DNA binding proteins that bind to 12 to 40 nucleotides long specific sequences. Modifying a meganuclease to bind to a new target sequence was complicated. There are approximately 200 synthetic meganucleases. Due to their inconvenience in designing and expense, they were rejected.
Zinc Fingers
These are the DNA binding proteins discovered in the year 2005. They recognize a specific DNA target sequence. Zinc fingers are composed of repeating units of triplet codons of 18 to 21 nucleotides. For efficient cutting, ZFN domains bind to a nuclease domain restriction endonuclease Fok1. Dimerization is necessary to activate Fok1 nuclease, ZFNs are used in pairs to recognize upstream and downstream of the target cut site.
TALENs
TALEN stands for Transcription activator-like effector nucleases derived from plant bacteria in the year 2011. Each TALEN repeats recognize a single base ( A, T, G, C) rather than a triplet codon. Four different TALEN domains are required to recognize any given DNA sequence.

The feasibility of different Genome editing tools (SOURCE)
CRISPR-Cas9
This technology was developed in the year 2012. CRISPR stands for Clustered Regularly Interspaced Short Palindromic Repeats. In this method, the task of DNA recognition is carried out by a piece of sgRNA (single-stranded guide RNA) that binds to a matching position in the DNA. It is a natural immune system found in bacteria in response to viral infections. The Cas 9 protein acts as a nuclease(molecular scissors) that cuts the recognized viral sequence. The CRISPR system uses RNA that can base pair with the complementary sequences of the viral genome which guides the Cas9 nuclease to cleave at correct sites. Another question raised is that how can a bacteria differentiate its genome fro the viral genome? The answer to this is the viral genome contains PAM sequences( Protospacer adjacent motifs) that allows bacteria to differentiate. The PAM sequence also directs Cas9 nuclease to cleave more accurately( usually 3 bases upstream of PAM sequence).
Applications of Nucleases
The first technique to be invented was based on molecules called zinc finger nucleases (ZFNs). ZFNs recognize bases in the DNA, bind to them, and cut the DNA near the binding site. Next, scientists figured out how to use another group of enzymes, called transcription activator-like effector nucleases (TALENs). Like ZFNs, TALENs can be used to cut DNA in specific places to introduce mutations. They recognize specific single nucleotides in the DNA. Both ZFN and TALEN are enzymes that recognize binding sites in the DNA and have been engineered to target various genomic regions. TALENs are easier to engineer because they bind to fewer bases in the DNA than ZFNs.
The Promise Of CRISPR Technology

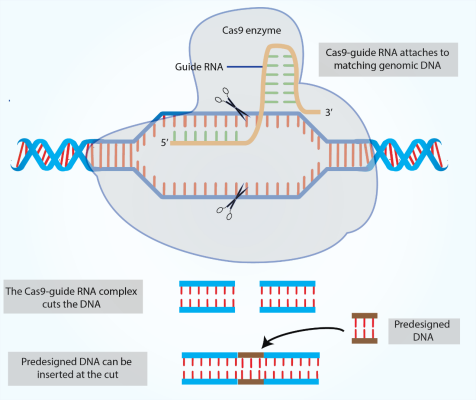
The Single-stranded Guide RNA(sg RNA) directs Cas9 protein to bind to the accurate cleaving site(SOURCE)
The so-called CRISPR/Cas9 method – short for Clustered Regularly Interspaced Short Palindromic Repeat/CRISPR Associated – is different. In this method, the task of DNA recognition is carried out by a piece of RNA that binds to a matching position in the DNA. Then the Cas9 nuclease cuts the DNA. As with ZFNs and TALENs, this process either “knocks out” the targeted gene or, if a matching piece of DNA has been added, the targeted gene mutates. It’s much easier and faster to engineer synthetic RNAs that fit targeted areas than it is to engineer enzymes like ZFNs or TALENs. Another advantage of CRISPR is that more than one gene can be targeted simultaneously by adding a variety of RNAs at once. This allows researchers to study diseases that are polygenic (caused by many genes) more effectively.

A cartoon indicating the action of the CRISPR-Cas9 system in bacterial immune system against viruses (SOURCE)
CRISPR is based on a molecular process that’s part of the bacterial immune system. Bacteria use it as a defense against viruses by cutting viral DNA/RNA into small pieces. The three steps of the bacterial immune CRISPR -Cas9 system
1. Adaptation/Memorization
2. Maturation and Assembly
3. Interference and Destruction
Several researchers, among them microbiologist Emmanuelle Charpentier, biochemist Jennifer Doudna, and neuroscientist/bioengineer Feng Zhang realized that this bacterial system could be adapted to change DNA in any cell and with high precision.
Applications Of CRISPR
CRISPR has taken labs worldwide by storm. Several companies, such as Editas Medicine and Intelia Therapeutics (Massachusetts) and CRISPR Therapeutics (Switzerland), are using CRISPR/Cas9 to find cures for diseases. Simultaneously, a big fight over patents for what some call “the biggest biotech discovery of the century” is going on.
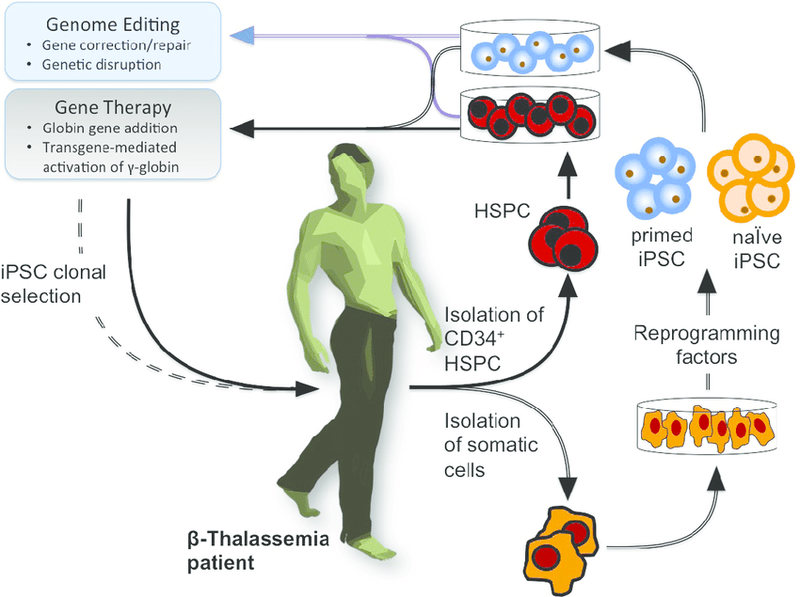
The role of Genome editing in treating Beta- Thalassemia(SOURCE)
Early in 2015 Chinese researchers published their attempts to modify the beta-thalassemia gene in a sample of 86 non-viable embryos from fertility clinics using CRISPR/Cas9. Only a small fraction of the embryos were successfully “fixed.” Scientists worldwide have called for a moratorium on genome editing methods of human cells until the methods have been safely assessed. The rapid changes and advancements in cloning technologies and genome editing methods make a public discussion about the ethics of human genetic engineering indispensable.
Researches
Experiment

In the paper, researchers led by Junjiu Huang, a gene-function researcher at Sun Yat-sen University in Guangzhou, tried to head off such concerns by using ‘non-viable’ embryos, which cannot result in a live birth, that were obtained from local fertility clinics. The team attempted to modify the gene responsible for β-thalassemia, a potentially fatal blood disorder, using a gene-editing technique known as CRISPR/Cas9. The researchers say that their results reveal serious obstacles to using the method in medical applications.
“I believe this is the first report of CRISPR/Cas9 applied to human pre-implantation embryos and as such the study is a landmark, as well as a cautionary tale,” says George Daley, a stem-cell biologist at Harvard Medical School in Boston, Massachusetts. “Their study should be a stern warning to any practitioner who thinks the technology is ready for testing to eradicate disease genes.”
gene editing in embryos could have a bright future because it could eradicate devastating genetic diseases before a baby is born. Others say that such work crosses an ethical line because the genetic changes to embryos, known as germline modification, are heritable, they could have an unpredictable effect on future generations. Researchers have also expressed concerns that any gene-editing research on human embryos could be a slippery slope towards unsafe or unethical uses of the technique.
Observations
The team injected 86 embryos and then waited 48 hours, enough time for the CRISPR/Cas9 system and the molecules that replace the missing DNA to act — and for the embryos to grow to about eight cells each. Of the 71 embryos that survived, 54 were genetically tested. This revealed that just 28 were successfully spliced and that only a fraction of those contained the replacement genetic material.
“If you want to do it in normal embryos, you need to be close to 100%,” Huang says.
Drawbacks

The team also found a surprising number of ‘off-target’ mutations assumed to be introduced by the CRISPR/Cas9 complex acting on other parts of the genome. This effect is one of the main safety concerns surrounding germline gene editing because these unintended mutations could be harmful. The rates of such mutations were much higher than those observed in gene-editing studies of mouse embryos or human adult cells. Huang notes that his team detected only a subset of the unintended mutations because their study looked only at a portion of the genome. “If we did the whole genome sequence, we would get many more,” he says. A high number of off-target mutations could be specific to the abnormal embryos used in the study. There are no examples of gene editing in normal embryos. The technique could operate differently in them.
Conclusion
The future aim is to decrease the number of off-target mutations using adult human cells or animal models. This may be done by tweaking the enzymes to guide them more precisely to the desired spot, introducing the enzymes in a different format that could help to regulate their lifespans and thus allow them to be shut down before mutations accumulate, or varying the concentrations of the introduced enzymes and repair molecules. He says that using other gene-editing techniques might also help. CRISPR/Cas9 is relatively efficient and easy to use, but another system called TALEN is known to cause fewer unintended mutations. The debate over human embryo editing is sure to continue for some time, however. CRISPR/Cas9 is known for its ease of use and more scientists will now start to work towards improving on Huang’s paper. This creates opportunities for scientists in any part of the world to do any kind of experiment.